Speed up Pandas in Python with Modin: A Comprehensive Guide
Updated on
Update (Nov 2025): Modin 0.32+ supports pandas 2.2, Ray 2.x, and Dask backends. Install with
pip install "modin[dask]"orpip install "modin[ray]"to select a backend, and keep pandas <3 pinned if your environment mixes extensions that have not declared pandas 3 support yet.
In the world of data science, the ability to manipulate and analyze large datasets is a crucial skill. One of the most popular libraries for data manipulation in Python is Pandas. However, as the volume of data grows, the performance of Pandas can become a bottleneck. This is where Modin comes in! In this comprehensive guide, we will explore how to speed up Pandas in Python with Modin and discuss its benefits, drawbacks, and best practices.
Want to quickly create Data Visualizations in Python?
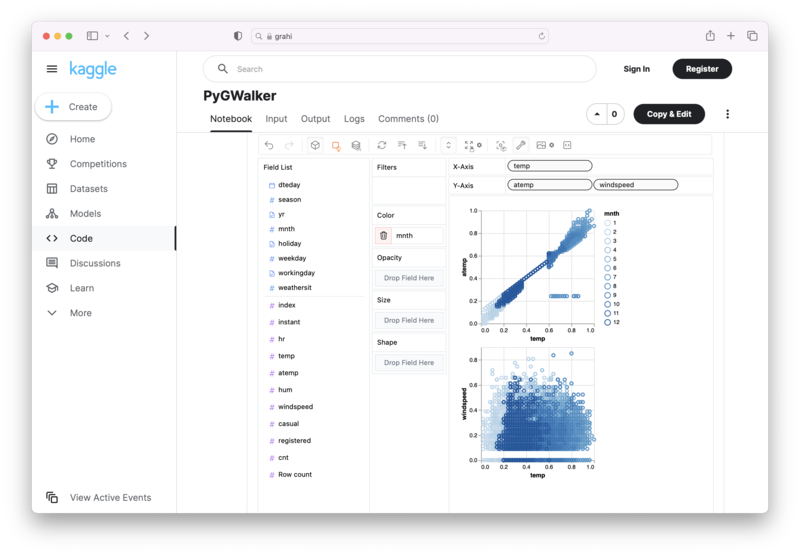
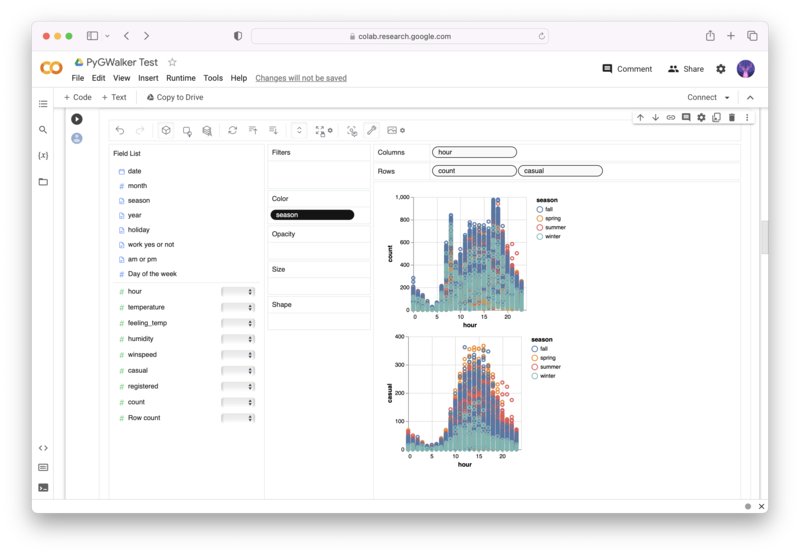
PyGWalker is an Open Source Python Project that can help speed up the data analysis and visualization workflow directly within a Jupyter Notebook-based environments.
PyGWalker (opens in a new tab) turns your Pandas Dataframe (or Polars Dataframe) into a visual UI where you can drag and drop variables to create graphs with ease. Simply use the following code:
pip install pygwalker
import pygwalker as pyg
gwalker = pyg.walk(df)You can run PyGWalker right now with these online notebooks:
And, don't forget to give us a ⭐️ on GitHub!
What is Pandas?
Pandas is an open-source library that provides data manipulation and analysis tools for Python. It offers data structures like Series and DataFrame, which are ideal for handling structured data. Pandas is known for its ease of use, flexibility, and powerful data manipulation capabilities.
What is Modin and how does it work?
Modin is a library designed to speed up Pandas by leveraging distributed computing and parallelization techniques. It is built on top of the Dask or Ray frameworks and aims to provide a more efficient and scalable solution for working with large data in Python. Modin works by dividing the DataFrame into smaller partitions and processing each partition in parallel, thereby accelerating the execution of Pandas operations.
How can Modin help in speeding up Pandas in Python?
Modin can significantly speed up Pandas operations by taking advantage of parallel computing and distributed processing. By partitioning the DataFrame and processing each partition concurrently, Modin can handle larger datasets and improve the performance of data manipulation tasks. Some benefits of using Modin to speed up Pandas in Python include:
-
Improved performance: Modin can provide a substantial performance boost for a wide range of Pandas operations, including filtering, sorting, and aggregation.
-
Ease of use: Modin offers a familiar API that is almost identical to that of Pandas, making it easy for users to adapt their existing code.
-
Scalability: Modin can handle larger datasets by distributing the computation across multiple cores or nodes in a cluster.
-
Flexibility: Modin supports various backends, such as Dask and Ray, allowing users to choose the most suitable framework for their specific use case.
Are there any disadvantages to using Modin?
While Modin offers several benefits for speeding up Pandas in Python, there are some potential drawbacks to consider:
-
Compatibility: Modin does not support all Pandas operations, which may require users to make adjustments to their existing code.
-
Overhead: Modin introduces additional overhead due to the partitioning and parallelization processes, which may impact performance for small datasets.
-
Dependencies: Modin relies on external frameworks like Dask and Ray, which may introduce additional complexity and dependencies to the project.
How can I install Modin in Python?
To install Modin, you can use the following command with pip:
pip install modinBy default, Modin uses the Dask backend. If you prefer to use the Ray backend, you can install it with the following command:
pip install modin[ray]Once installed, you can use Modin just like you would use Pandas. Simply replace the import pandas as pd statement with import modin.pandas as pd, and the rest of your code should remain unchanged. For example:
import modin.pandas as pd
data = pd.read_csv("large_dataset.csv")
filtered_data = data[data["column_name"] > 100]Modin vs. Pandas: Which one is faster?
In most cases, Modin is faster than Pandas when working with large datasets. Modin's parallelization and distributed computing capabilities enable it to process data more efficiently, reducing the time required for common operations. However, for smaller datasets, the performance difference may be negligible or even slightly worse due to the overhead introduced by Modin.
What are the alternatives to Modin for speeding up Pandas?
While Modin is an excellent option for speeding up Pandas in Python, there are alternative libraries and techniques to consider:
-
Dask: Dask is a parallel computing library that can be used directly to speed up Pandas operations by distributing them across multiple cores or nodes. Dask provides a familiar API that closely resembles Pandas, making it a good choice for users looking for a more granular level of control over parallelization.
-
Vaex: Vaex is a high-performance library that enables efficient data manipulation and visualization for large datasets. It uses a lazy evaluation approach, which means that operations are not executed immediately but rather deferred until the results are needed, helping to reduce memory usage and improve performance.
-
Optimizing Pandas: You can also optimize your Pandas code by using vectorized operations, efficient data types, and other performance-enhancing techniques.
How to handle big data with Pandas in Python?
Handling big data with Pandas in Python can be challenging due to performance limitations. However, there are several strategies you can employ to work with large datasets effectively:
- Use libraries like Modin or Dask to leverage parallel computing and distributed processing.
- Optimize your Pandas code to take advantage of vectorized operations and efficient data types.
- Break down your dataset into smaller chunks and process them one at a time.
- Use a data storage solution like Apache Arrow or Parquet to store and load your data more efficiently.
- Consider using other big data processing frameworks like Apache Spark for more complex and large-scale data manipulation tasks.
What are the best practices for working with Pandas in Python?
To ensure optimal performance and ease of use when working with Pandas in Python, consider the following best practices:
- Use vectorized operations to perform element-wise calculations on entire columns or DataFrames.
- Choose appropriate data types to minimize memory usage and improve performance.
- Use the
inplaceparameter when possible to modify DataFrames without creating new objects. - Opt for built-in Pandas functions instead of custom Python functions for improved performance.
- When working with large datasets, consider using libraries like Modin or Dask to improve performance through parallelization and distributed computing.
Conclusion
By following these best practices and leveraging the power of Modin, you can speed up your Pandas dataframes in Python, making it easier to handle big data and optimize your data processing workflows.
More Pandas Tutorials: